Proposals for our CSP New Investigator call are due October 4, 2024
Proposals for our CSP New Investigator call are due October 4, 2024
Read our JGI Data Policy
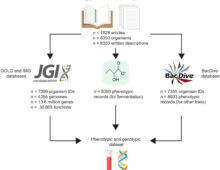
Harnessing BER-funded data resources, researchers built an interactive and comprehensive database of fermentative prokaryotes. The Science Most organisms use oxygen to convert food into energy. However, in environments with little or no oxygen, life had found other ways to produce energy, … [Read More]